Rebuilding small GTPase networks in vitro
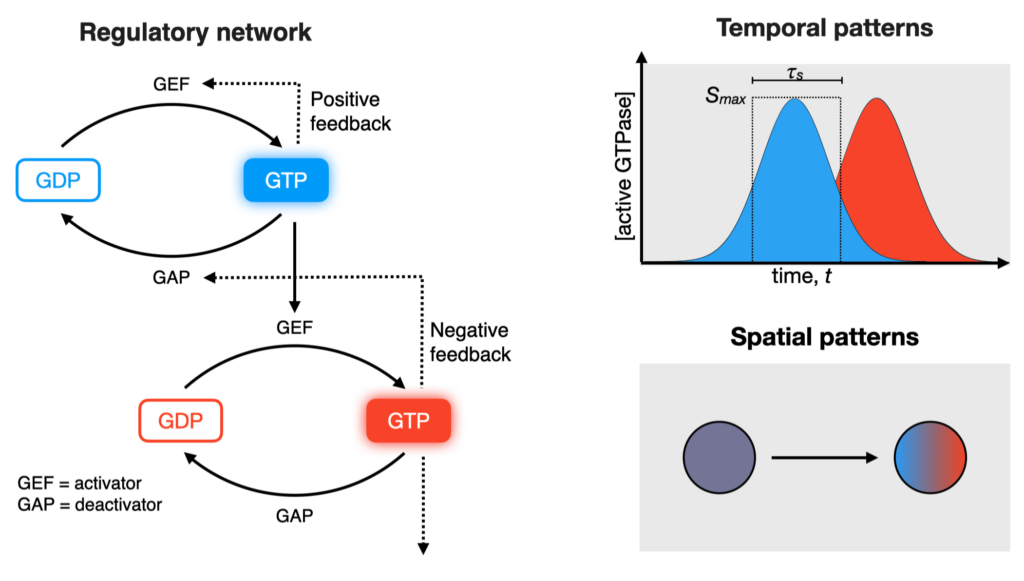
Small GTPases are molecular switches that can exist in an ON- and OFF-state and thereby regulate a wide variety of cellular processes in eukaryotic cells, such as vesicle formation, transport and fusion. The activities of these proteins are regulated in complex regulatory networks, which can give rise to dynamic activity patterns in the living cells.
We are using an in vitro reconstitution approach to understand the logic and mechanisms of the regulatory networks that control small GTPases.
Recently, we have used our in vitro reconstitution approach to answer the following questions:
- What gives rise to collective activation of Rab5 in space and time ?
While the GTPase switch is well understood on a molecular level, cellular functions of small GTPases usually rely on their collective behavior, i.e. a large number of proteins need switch simultaneously at the right time and place. How this collective switching is achieved is often not known. In the case of the small GTPase Rab5, previous experimental and theoretical evidence suggested that it could be the result of a complex biochemical circuits that includes a positive feedback. However, an experimental verification of this hypothesis had been missing.
In this collaborative project with the group of Timothy Saunders, we combined in vitro reconstitution experiments with theoretical modeling to study the emergent properties of a Rab5 GTPase activation network comprised of Rab5, its GDP dissociation inhibitor (GDI) and nucleotide exchange factor Rabex5, the scaffold protein Rabaptin5 and biomimetic membranes.
We found that collective GTPase switching relies on a positive feedback and a involves mechanism that repress indiscriminate noise amplification. Notably, we demonstrate that collective switching can spread on the membrane surface as a traveling wave of Rab5 activation.
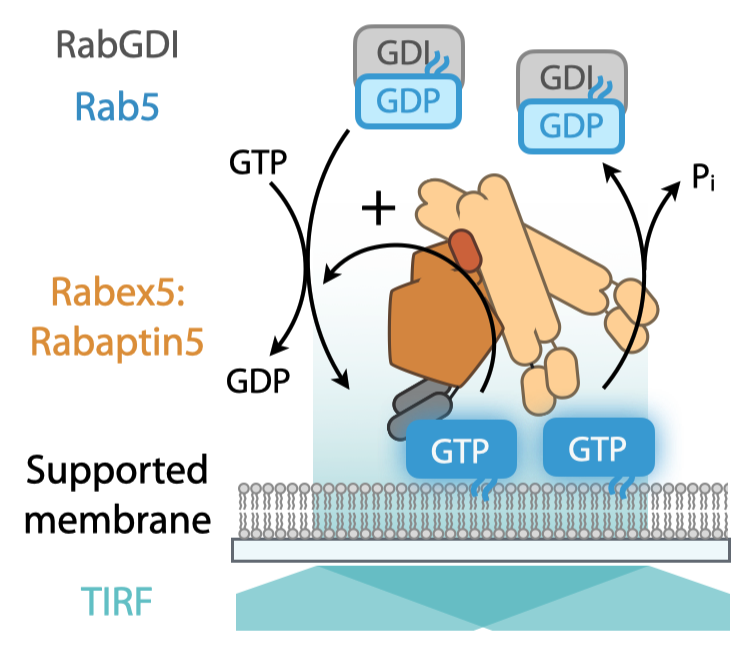

The Rab5 activation network gives rise to bistability and the coexistence of the ON- and OFF-state. As a result we can observe traveling waves of Rab5 activation on the membrane. Importantly, such self-organized spatial patterns share features with activity patterns found for Rab5 in vivo.
Together, these findings reveal how biochemical networks control vectorial vesicle trafficking pathways and how their non-equilibrium properties define the spatiotemporal organization of the cell (Bezeljak et al., 2020).
- What controls the activity of Arf6 in a nascent neuron?
Eukaryotic cell differentiation depends on a precise control of their internal organization, which is the result of a complex dynamic interplay between the cytoskeleton, molecular motors, signaling molecules and membranes.
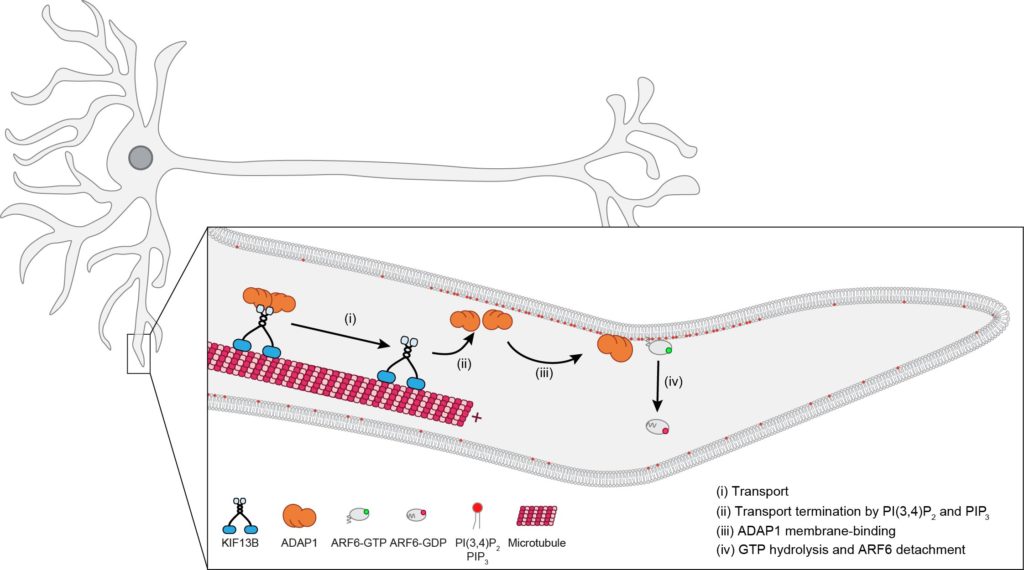
For example, in the developing neuron the protein ADAP1 has been implicated in the control of the small GTPase Arf6 and dendrite branching. It is also thought to mediate axon specific delivery of PI(3,4,5)P3 together with the motor protein KIF13B, thus contributing to phosphoinositide polarity (Hansen et al., 2017). However, how these different roles of ADAP1 are coordinated remains unclear.
We studied ADAP1’s functions using in vitro reconstitutions and found that KIF13B transports ADAP1 along microtubules. Instead of being a cargo, as previously believed, we found that PIP3 instead acts as a stop signal for this transport. At the same time, PIP3 and PI(3,4)P2 activate ADAP1s enzymatic activity to catalyze GTP hydrolysis by Arf6. Our results support a model, by which KIF13B transports ADAP1 towards high concentrations of PIP3/PI(3,4)P2 in the plasma membrane, where it disassociates from the motor and inactivates Arf6. Importantly, these findings demonstrate that phosphoinositides can act as cargo-release factors and challenge current models for how polarity is established in neurons (Duellberg & Auer et al., 2021).
PEOPLE WORKING ON THIS PROJECT
Albert, Ivana, Arsenii, Anita
Collaborators
- Carrie Bernicky (ISTA, Klosterneuburg, Austria)
- Florian Schur (ISTA, Klosterneuburg, Austria)
- Timothy Saunders (University of Warwick, UK)
RELEVANT PUBLICATIONS
Martin Loose, Albert Auer, Gabriel Brognara, Hanifatul Rahmah Budiman, Lukasz Kowalski, Ivana Matijević
In vitro reconstitution of small GTPase regulation (Review)
Febs Lett (2023), vol. 597, 762–777.
Christian Duellberg, Albert Auer, Nicola Canigova, Katrin Loibl, Martin Loose.
In vitro reconstitution reveals phosphoinositides as cargo-release factors and activators of the ARF6 GAP ADAP1
PNAS (2021), vol. 118 (1) e2010054118.
U. Bezeljak, H. Loya, B. Kaczmarek, T. E. Saunders and M. Loose.
Stochastic activation and bistability in a Rab GTPase regulatory network.
PNAS (2020), vol. 117(12):6540-6549.
P. A. Nguyen*, A. C. Groen*, M. Loose, K. Ishihara, M. Wühr, C. M. Field, and T. J. Mitchison.
Spatial Organization of Cytokinesis Signaling Reconstituted in a Cell-Free System.
Science (2014) vol. 346, 244-247
FUNDING

This research project (“GTPaseNET”) has received funding from the European Research Council (ERC), Grant number: 101045340.
